DNA repair
FANCJ helicase unfolds DNA-protein cross-links
05.01.2023
Endogenous and exogenous agents generate DNA-protein cross-links (DPCs), whose replication-dependent degradation by the SPRTN protease suppresses aging and liver cancer. SPRTN is activated after the replicative CMG helicase bypasses a DPC and polymerase extends the nascent strand to the adduct.
In a study published in Molecular Cell, Julian Stingele and his team, in collaboration with the lab of Johannes Walter, identified a role for the 5’ to 3’ helicase FANCJ in DPC repair. In addition to supporting CMG bypass, FANCJ is essential for SPRTN activation. FANCJ binds ssDNA downstream of the DPC and uses its ATPase activity to unfold the protein adduct, which exposes the underlying DNA and enables cleavage of the adduct. FANCJ-dependent DPC unfolding is also essential for translesion DNA synthesis past DPCs that cannot be degraded. In summary, their results show that helicase-mediated protein unfolding enables multiple events in DPC repair.
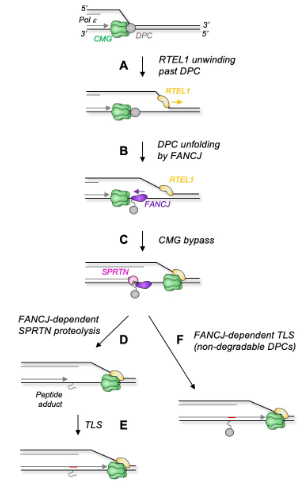
Figure: Model for FANCJ's role in DPC repair
(A) Upon replisome collision with a leading strand DPC, RTEL1 (and possibly FANCJ) translocates along the undamaged lagging strand template, exposing ssDNA beyond 30 the DPC, which supports CMG bypass via an unknown mechanism. TRAIP ubiquitylates the DPC before CMG bypass occurs. (B) FANCJ loads onto the singlestranded leading strand template downstream of the DPC and translocates back towards the DPC, which it remodels. (C) CMG bypasses the DPC. (D) After bypass, the DPC undergoes proteolysis by the proteasome or SPRTN, whose activity depends on FANCJ-dependent DPC unfolding. (E) Finally, the leading strand is extended past the peptide. If the DPC fails to be degraded, FANCJ-dependent DPC unfolding enables TLS past the intact adduct (F). Source: Molecular Cell 2023.
Original Publication:
The FANCJ Helicase Unfolds DNA-Protein Cross-Links to Promote Their Repair
Yaneva D, Sparks JL, Donsbach M, ZhaoS, Weickert P, Bezalel-Buch R, Stingele J, and Walter JC
Mol Cell. 2023, Jan 5;83(1):43-56.e10. https://doi.org/10.1016/j.molcel.2022.12.005

