Structural Biology
Enigmatic nuclease diversity of Mre11-Rad50 elucidated
19.08.2022
Hopfner lab reveals cryoEM structures of the Mre11-Rad50 complex and the structural mechanism of processing blocked DNA ends and hairpins
DNA double-strand breaks (DSBs) threaten genome stability and are linked to tumorigenesis in humans. Repair of DSBs requires the removal of attached proteins and hairpins through a poorly understood but physiologically critical endonuclease activity by the Mre11-Rad50 complex. The Hopfner group report cryoelectron microscopy (cryo-EM) structures of the bacterial Mre11-Rad50 homolog SbcCD bound to a protein-blocked DNA end and a DNA hairpin. The structures reveal that Mre11-Rad50 bends internal DNA for endonucleolytic cleavage and show how internal DNA, DNA ends, and hairpins are processed through a similar ATP-regulated conformational state. Furthermore, Mre11-Rad50 is loaded onto blocked DNA ends with Mre11 pointing away from the block, explaining the distinct biochemistries of 3′ → 5′ exonucleolytic and endonucleolytic incision through the way Mre11-Rad50 interacts with diverse DNA ends. In summary, the results unify Mre11-Rad50’s enigmatic nuclease diversity within a single structural framework and reveal how blocked DNA ends and hairpins are processed.
The results also provide an interesting approach for further studies explains first author Fabian Gut: "An unexpected observation made was the DNA bending by Mre11-Rad50 in the endonuclease state. This provides space and ideas for future studies, focusing on the physiological relevance of DNA mechanical or local sequence-specific properties."
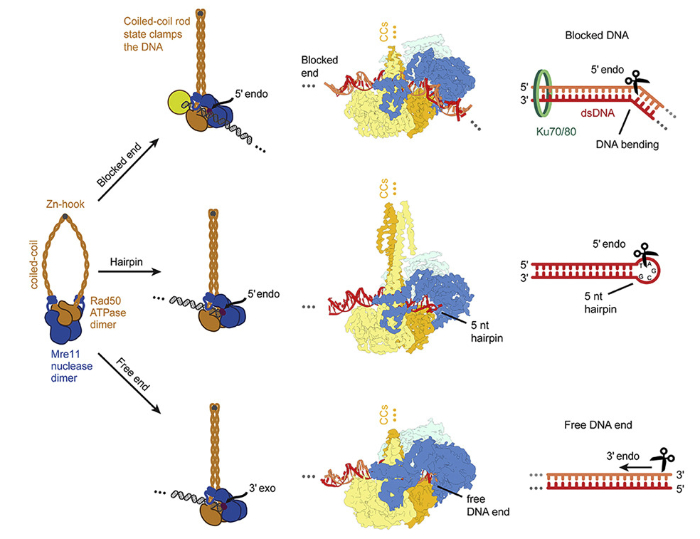
A unified mechanistic basis for Mre11-Rad50's endo-, exonuclease and hairpin-opening activities, revealing how MR processes a variety of obstructed DNA double-strand break ends. Image: Hopfner group
Original Publication:
Structural mechanism of endonucleolytic processing of blocked DNA ends and hairpins by Mre11-Rad50.
Gut F, Käshammer L, Lammens K, Bartho JD, Boggusch AM, van de Logt E, Kessler B, Hopfner KP.
Mol Cell. 2022 Aug 12:S1097-2765(22)00715-8. doi: 10.1016/j.molcel.2022.07.019.

