Structural Biology: Ribosomes
Structural basis for clearing of ribosome collisions
17.02.2023
Beckmann group in collaboration with Inada lab unveils the structural basis of the ribosome-associated quality control trigger complex (RQT) pathway, which is activated when ribosomes collide during persistent stalling. The RQT complex splits the stalled ribosome, thus clearing harmful traffic jams in the cell.
Abstract
Translation of aberrant messenger RNAs can cause stalling of ribosomes resulting in ribosomal collisions. Collided ribosomes are specifically recognized to initiate stress responses and quality control pathways. Ribosome-associated quality control facilitates the degradation of incomplete translation products and requires dissociation of the stalled ribosomes. A central event is therefore the splitting of collided ribosomes by the ribosome quality control trigger complex, RQT, by an unknown mechanism. Here we show that RQT requires accessible mRNA and the presence of a neighboring ribosome. Cryogenic electron microscopy of RQT-ribosome complexes reveals that RQT engages the 40S subunit of the lead ribosome and can switch between two conformations. We propose that the Ski2-like helicase 1 (Slh1) subunit of RQT applies a pulling force on the mRNA, causing destabilizing conformational changes of the small ribosomal subunit, ultimately resulting in subunit dissociation. Our findings provide conceptual framework for a helicase-driven ribosomal splitting mechanism.
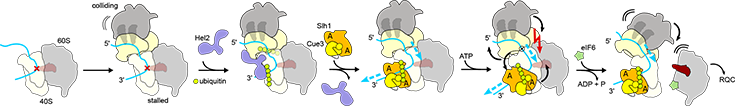
After stalling and subsequent collision of trailing ribosomes, Hel2 recognizes the collision interface and (poly)ubiquitinates uS10. RQT binds polyubiquitinated stalled ribosomes and engages with accessible 3′-mRNA emerging from the stalled lead ribosome. A pulling force on the mRNA (blue arrows) may cause head-swiveling, destabilization of the lead ribosome and ultimately drive the trailing ribosome like a wedge between the subunits of the lead ribosome. The red cross indicates a stall on problematic mRNA (blue); P-site tRNA in the stalled ribosome tRNA is indicated in transparent red and hybrid tRNAs in the collided ribosomes are transparent gray; Green circles indicate ubiquitin; black arrows indicate possible directions of movement and black curved lines indicate movement/flexibility; A = ATP-binding site. Source: Nat. Commun. 14(1):921, 2023
Original publication:
Structural basis for clearing of ribosome collisions by the RQT complex
Best K, Ikeuchi K, Kater L, Best D, Musial J, Matsuo Y, Berninghausen O, Becker T, Inada T, Beckmann R.
Nat Commun. 2023 Feb 17;14(1):921. doi: 10.1038/s41467-023-36230-8.

