Halic Lab - Research
- Research
- Heterochromatin Formation
Regulation of genome expression is essential for many cellular processes including cell proliferation, differentiation, development and viability. It has become clear over the past decade that small RNA silencing pathways are involved in the cellular control of gene expression and in protection of the genome against mobile repetitive DNA sequences, retroelements and transposons. Small RNAs interact with target RNAs by base pairing interactions and promote chromatin modifications, translational inhibition or degradation of complementary RNAs to induce gene silencing.
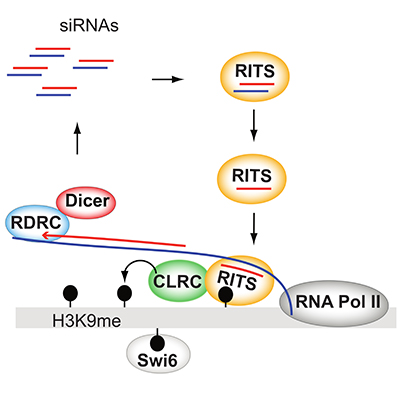 In the fission yeast Schizosaccharomyces pombe, small RNA pathways act at the chromatin or DNA level and are essential for formation of heterochromatin at centromeric repeats (see Figure). Small RNA mediated heterochromatin formation is required for deposition of centromere specific nucleosomes and for proper chromosome segregation. Small interfering RNAs (siRNAs), the mediator of centromeric silencing, are generated by enzyme Dicer from double-stranded RNA (dsRNA) and subsequently loaded onto Argonaute protein. siRNAs direct the inactivation of target RNAs by guiding the Argonaute containing RNA Induced Transcriptional Silencing (RITS) complex to complementary centromeric target sequences. This leads to the recruitment of RNA dependent RNA polymerase complex (RDRC) to centromeric transcripts, synthesis of new dsRNAs and siRNA amplification (see Figure). In addition to RDRC, RITS complex recruits the histone 3 lysine 9 (H3K9) methyltransferase complex CLRC to chromatin, which leads to repressive H3K9 methylation, subsequent recruitment of fission yeast heterochromatin protein HP1 and heterochromatin formation (see Figure).
In the fission yeast Schizosaccharomyces pombe, small RNA pathways act at the chromatin or DNA level and are essential for formation of heterochromatin at centromeric repeats (see Figure). Small RNA mediated heterochromatin formation is required for deposition of centromere specific nucleosomes and for proper chromosome segregation. Small interfering RNAs (siRNAs), the mediator of centromeric silencing, are generated by enzyme Dicer from double-stranded RNA (dsRNA) and subsequently loaded onto Argonaute protein. siRNAs direct the inactivation of target RNAs by guiding the Argonaute containing RNA Induced Transcriptional Silencing (RITS) complex to complementary centromeric target sequences. This leads to the recruitment of RNA dependent RNA polymerase complex (RDRC) to centromeric transcripts, synthesis of new dsRNAs and siRNA amplification (see Figure). In addition to RDRC, RITS complex recruits the histone 3 lysine 9 (H3K9) methyltransferase complex CLRC to chromatin, which leads to repressive H3K9 methylation, subsequent recruitment of fission yeast heterochromatin protein HP1 and heterochromatin formation (see Figure).
In the nascent transcript cycle described above, the RNAi machinery localizes to chromatin-bound transcripts and mediates their processing into siRNAs, which promote heterochromatin assembly and the generation of additional siRNAs. However, it is unclear how siRNA generation and heterochromatin assembly are initiated in the first place, since the cycle begins with an siRNA-programmed RITS.

