Canzar Lab - Algorithmic Bioinformatics
 Research Topics
Research Topics
- Computational Genomics & Transcriptomics
- Algorithm Engineering
- Single Cell Genomics
- Combinatorial Optimization
Part of the group relocated to Universität Regensburg
Dr. Stefan Canzar
canzar@genzentrum.lmu.de or
stefan.canzar@informatik.uni-regensburg.de
Next-generation sequencing instruments produce a huge number of short DNA 'reads', each of which carries little information by itself. These reads therefore have to be pieced together by well-engineered algorithms to reconstruct biologically meaningful measurements. The lab's goal is the development of accurate mathematical models, efficient algorithms, and usable software to solve these complex, high-dimensional puzzles. Read more...
Selected Publications
Partitioning RNAs by length improves transcriptome reconstruction from short-read RNA-seq data.
Ringeling FR, Chakraborty S, Vissers C, Reiman D, Patel AM, Lee KH, Hong A, Park CW, Reska T, Gagneur J, Chang H, Spletter ML, Yoon KJ, Ming GL, Song H, Canzar S
Nature Biotechnology. 2022. doi:10.1038/s41587-021-01136-7
A generalization of t-SNE and UMAP to single-cell multimodal omics.
Van Do H, Canzar S.
Genome Biology. 2021;22(1):130. doi: 10.1186/s13059-021-02356-5.
Linear-time cluster ensembles of large-scale single-cell RNA-seq and multimodal data.
Van Do H, Rojas Ringeling F, Canzar S.
Genome Research. 2021;31(4):677-688 doi: 10.1101/gr.267906.120.
Sphetcher: Spherical thresholding improves sketching of single-cell transcriptomic heterogeneity.
Van Do H, Elbassioni K, Canzar S.
iScience. 2020;23(6):101126. doi: 10.1016/j.isci.2020.101126.
More publications
News
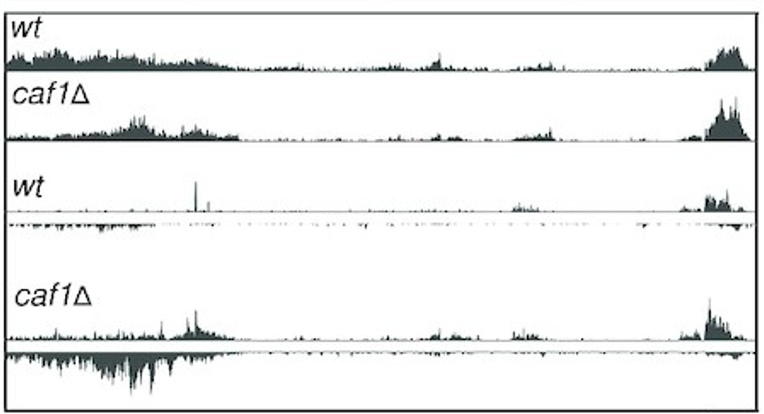
30.05.2022 New paper
Ccr4–Not complex reduces transcription efficiency in heterochromatin. Congrats Pablo!
10.01.2022 Ladder-seq is out!
Partitioning RNAs by length improves transcriptome reconstruction from short-read RNA-seq data.
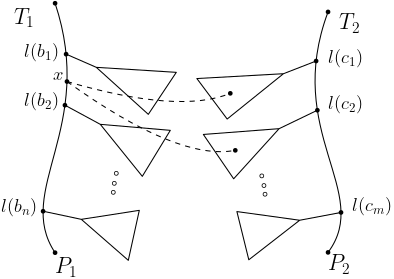
20.07.2021 - New preprint
Anti Tai Mapping for Unordered Labeled Trees
01.07.2021 - DFG funding
We are part of LETSIMMUN, a collaborative research center funded by the DFG
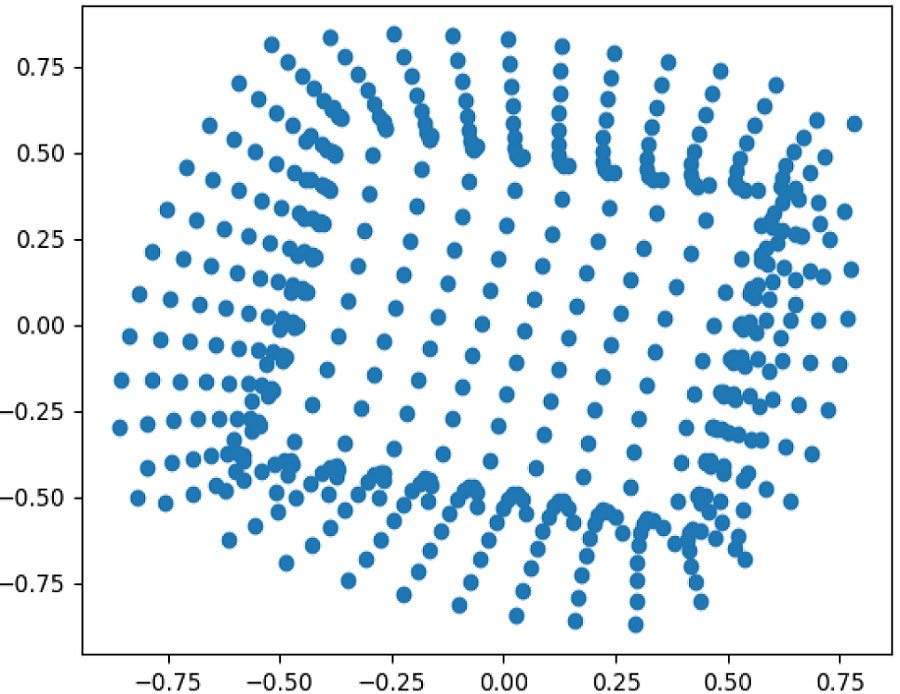
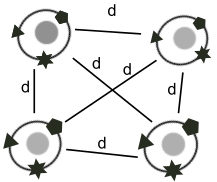
27.06.2021 - New preprint
Fast metric multidimensional scaling using a neural network
03.05.2021 - New paper
j-SNE and j-UMAP generalize t-SNE and UMAP to single-cell multimodal omics
19.02.2021 - New paper
Specter for (fast) multi-modal clustering of single cells accepted at Genome Research.
30.01.2021 - New paper
McSplicer infers a probabilistic model of alternative splicing
21.02.2020 - New paper
Single-cell sampling by Van Hoan Do accepted for presentation at RECOMB-seq and for publication at iScience
18.12.2019 - DAAD Project funding
Application of Generic Optimizer in Medical Imaging Classification Problems