Beckmann Lab - Research
- Research
- mRNA Surveillance
- Membrane Protein Biogenesis
- Ribosome Biogenesis
Quality Control and mRNA Surveillance on the Ribosome
During translation a functional protein is made from mRNA. If, however, this mRNA is faulty, the ribosome is slowed down or even paused, which in turn leads to the induction one of several quality control pathways to resolve this situation. The defective mRNA as well as the not fully translated protein needs to be degraded and the stalled ribosome recycled. Here, the recognition of a stalled ribosome turned out to be the committed step for initiation of all downstream quality control pathways.
mRNA can be faulty due to multiple reasons each calling for its own specific mRNA surveillance pathway. Nonsense-mediated decay (NMD) is caused by premature stop-codons, whereas the lack of a stop codon, truncated or damaged mRNA as well as stable secondary structure can cause pathways called non-stop decay (NSD) and no-go decay (NGD), which all have in common the slowed down or stalled ribosome as a central hub to initiate the degradation pathways. During this the slowed down ribosome is recognized, decay factors for the faulty mRNA and the truncated protein are recruited and the ribosome is recycled. Recent studies showed that ubiquitinylation of ribosomal proteins follow ribosome stalling and turn out to be crucial for the degradation of the aberrant peptide in the ribosome quality control (RQC) pathway. Furthermore, a number of helicases (e.g. Ski2, Slh1, Dhh1 or Upf1) play a role in triggering molecular events downstream of stalled ribosome recognition.
The goal is to understand the structural basis of these surveillance processes and to disentangle the interplay between the ribosome, the mRNA, the nascent peptide and the numerous factors involved.
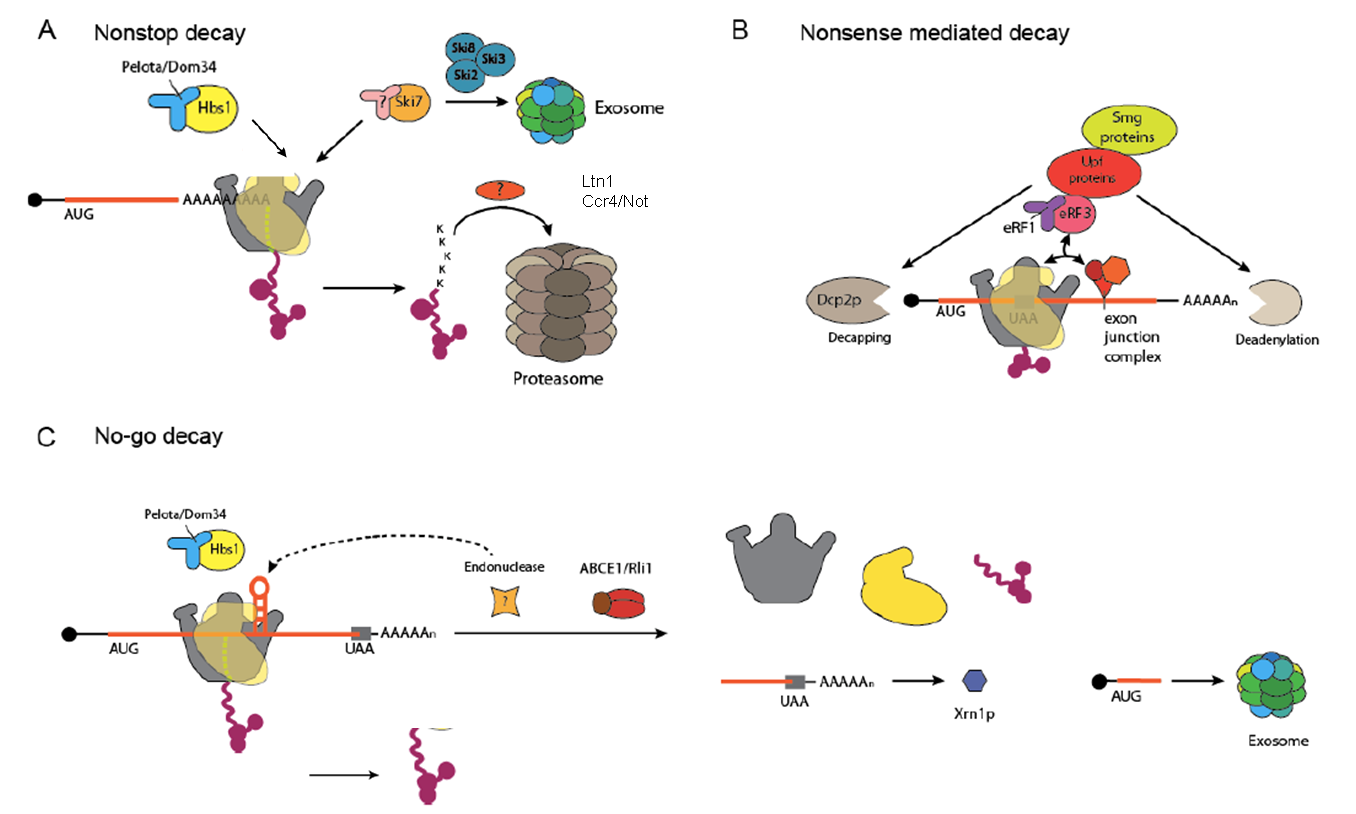
Overview of eukaryotic mRNA surveillance pathways (created by Sibylle Franckenberg).

