Förstemann Lab - Research
- Research
- Genome Surveillance
- siRNA Biogenesis
- Genome Engineering Tools
How is double-stranded RNA generated at specific sites in the genome of somatic cells?
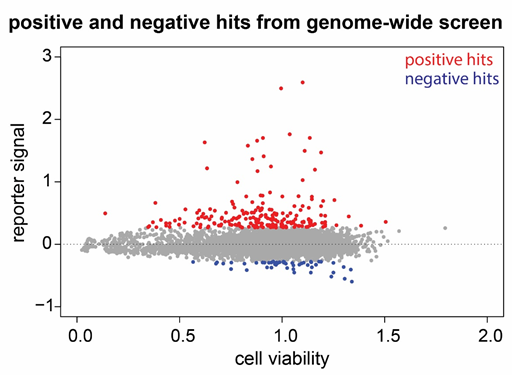
This is a central question to understand how the endo-siRNA system can fend off excessive transposable element activity. Normal transcription only produces RNA corresponding to one strand of the genome. Yet, to make siRNAs double-stranded RNA is needed. The decision to convergently transcribe a given region of DNA into sense and antisense RNA (giving rise to dsRNA) is thus a self vs. non-self distinction task. This is mechanistically related to siRNA biogenesis at a DNA double-strand break. The latter depends on active transcription and we have carried out a genome-wide screen in cultured cells to identify the underlying mechanism(s). For example, we found out that stalled splicing of the affected transcript is important to identify aberrant RNA molecules.