Blum Lab - Genomics
1) SARS-CoV-2 sequencing
SARS-CoV-2 phylogeny and frequencies of the currently 10.000 genomes sequenced in our laboratory within the Bay-VOC project. The samples are provided by the group of AG Keppler and AG Protzer (TUM) and sequenced in a weekly manner for the SARS-CoV-2 surveillance in Bavaria.
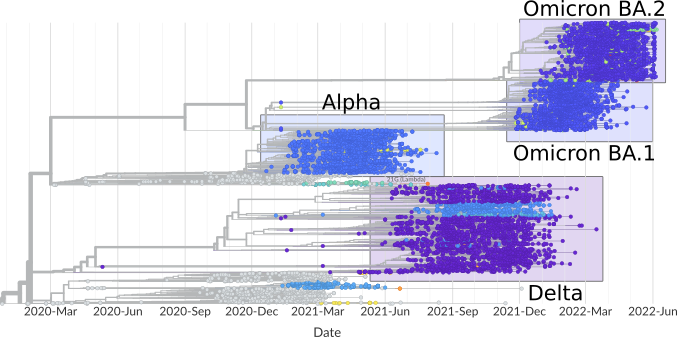

Phylogenetic tree of sequenced SARS-CoV-2 genomes.
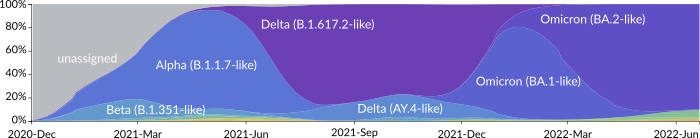
Frequencies of SARS-CoV-2 variants showing the waves of variants of concern over time.
Continuously updated data are availble on the Bay-VOC webpage.
2) Transcriptomics of early embryonic development
The early embryonic development is accompanied by complex processes like degradation of maternal RNAs or proteins stored in the oocytes. Next generation sequencing enables capturing transcripts down to single copies and thus provides unprecedented insights into developmental processes. We use sensitive methods for sequencing library construction like single primer isothermal amplification for detailed analyses of embryonic transcriptomes. Thus we studied transcripts that are not present in oocytes or are transcribed allele specific or provide access to de novo synthesis of RNA by detection of intronic sequences in primary transcripts.
Read more: Graf et al., Proc Natl Acad Sci U S A. 2014 Mar 18; 111(11):4139-44.
3) Epidemiology of Mycobacterium caprae
During the last decade Mycobacterium caprae infections have been frequently reported in cattle and red deer along the Bavarian and Austrian Alps. Several hundred M. caprae isolates recovered from infected animals provide an ideal source for molecular epidemiology by whole genome sequencing. Since available genomic DNA is heat denatured due to safety regulations, we had to adapt library protocols to enable whole genome sequencing of these samples. We use genomic data to study e.g. molecular evolution of geographically restricted substrains and the transmission dynamics of M. caprae within cattle herds.
Read more: Bröckl et al., Vet Res Commun. 2017 Jun;41(2):113-128.
4) Whole animal transcriptomics of disease models
The ultimate way to study the holistic impact of a disease like diabetes mellitus is to perform multi-omics analyses of all tissues of an affected organism and to systematically study influence on organ crosstalk.
We have access to random systematic sampled tissues from the Munich MIDY Pig Biobank modelling the poorly controlled diabetes mellitus in humans. We contribute sophisticated transcriptome analyses by high throughput sequencing. Using miniaturization and automation hundreds of sequencing libraries will be constructed and sequenced. Analyses of sequencing data and output of other -omics technologies such as targeted metabolomics, lipidomics and proteomics will pave the way to unprecedented insights into organ crosstalk in diabetes mellitus.
Read more: Blutke et al., Mol Metab. 2017 Aug; 6(8): 931–940.

